By altering the technical design of polymerase chain reaction (PCR) tests for malaria drug resistance, it's possible to build a simpler, faster assay suitable for testing whole blood in the field, particularly in low-resource settings, Vanderbilt University researchers reported June 13 in a proof-of-principle study published online in the Journal of Molecular Diagnostics.
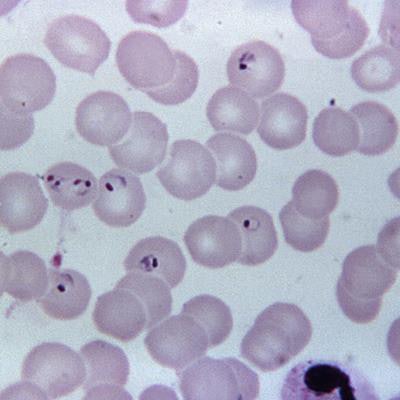
Co-lead investigator Mindy Leelawong, PhD, and colleagues reported that they have validated a point-of-care platform for a field-deployable PCR instrument, based on a study showing the test can differentiate between the wild-type form of the malaria-causing parasite Plasmodium falciparum and mutant types that are resistant to chloroquine, an older treatment for the disease.
"Our strategy greatly simplifies single-nucleotide polymorphism detection and provides a more accessible alternative for antimalarial resistance surveillance in the field," Leelawong et al wrote.
The study establishes the principle of a new approach, but it may take years for the test to be validated and broadly available. The most likely use for the technology would be in low-resource settings, said Frederick Haselton, PhD, who has patented the PCR instrument but not the underlying molecular biology tools, along with Nicholas Adams, PhD, both co-authors. These types of assays are usually vetted by the World Health Organization (WHO) and then rolled out on a country-by-country basis, Haselton explained in an email to LabPulse.com.
Confronting resistance, preventing disease spread
The assay was developed in response to unmet need for a test to evaluate resistance to treatments in low-resource settings. Years ago, the researchers noted, organisms developed resistance to the malaria treatments chloroquine and sulfadoxine-pyrimethamine, and a similar pattern is now being reported for the newer treatment artemisinin in some countries.
Monitoring resistance is important to prevent further spread of disease, and this is achieved through the detection of single-nucleotide polymorphisms (SNPs) using molecular techniques such as sequencing or real-time PCR.
"However, the tests are relatively expensive, are labor intensive, and are generally performed in central laboratories," Leelawong et al wrote. "We sought to make SNP genotyping for antimalarial drug resistance more accessible outside of central laboratories by developing an assay that has the potential to be performed directly on blood in a point-of-care setting."
In the U.S., the Centers for Disease Control and Prevention (CDC) recommends that all diagnosed cases of malaria be evaluated for drug resistance at its own central malaria testing laboratory, starting with PCR testing to confirm the species of parasite that caused the disease.
Research funded by NIH
The Vanderbilt University study was supported by a grant from the U.S. National Institutes of Health (NIH) Small Business Technology Transfer program and the NIH-funded Vanderbilt-Zambia Network for Innovation in Global Health Technologies.
An important feature of the new approach is that the probes in the PCR instrument were modified to be compatible with the optical properties of blood. In addition, locked nucleic acids (LNAs) were incorporated into the probes. A single drop of blood from a finger prick is tested with a small portable device.
"The LNA probes designed for this study enable SNP genotyping directly from blood," the investigators wrote. "As a consequence, drug-resistance testing for malaria can be performed without an additional DNA extraction step, which significantly decreases the test time and complexity."
The change provides a "critical advantage for SNP genotyping" that has been utilized for several tests, including chlamydia genotyping, the authors noted. The mutation associated with chloroquine resistance is now present in some circulating parasites in some regions, and this particular study was meant to serve as a proof of principle for testing resistance to other treatments. A future goal is to adapt the technology to look at mutations associated with resistance to artemisinin, the drug currently recommended by the WHO for first-line treatment of malaria, Leelawong told LabPulse.com.