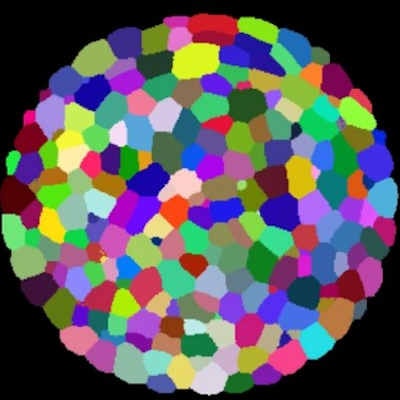
Researchers have developed a novel 3D cell segmentation framework called 3DCellSeg based on deep learning, according to an article published in Scientific Reports on January 10. The model can serve as a powerful tool to detect diseases in cells, including cancer.
"Our results suggest that 3DCellSeg can serve a powerful biomedical and clinical tool, such as histo-pathological image analysis, for cancer diagnosis and grading," wrote the authors, led by Andong Wang, PhD, of the department of electrical and electronic engineering at the University of Hong Kong.
The challenge of cell segmentation
An important prerequisite for biological image processing is cell segmentation, which involves identifying cells as objects of an image and tracking those objects from one image to the next. However, accurately segmenting densely packed cells in 3D images of cell membranes remains a challenge.
Existing approaches to cell segmentation are based on machine learning, in particular deep learning using convolutional neural networks (CNNs). However, such systems are difficult to reuse because they require the manual setting of "hyperparameters" (parameters used to control the learning process) on new datasets.
A second challenge is that once a segmentation system is up and running, it typically has low accuracy in regions with densely packed ("clumped") cells, where cells in the foreground occlude those in the background, making it difficult to identify different cell instances.
Meeting the challenge
To address these challenges, the authors developed a model called 3DCellSeg with a novel two-stage processing pipeline.
The first stage is semantic segmentation, where the input is a 3D cell membrane image and the output consists of three masks, indicating whether a voxel is the cell foreground, membrane, or background, respectively. This stage uses a separate loss function specifically geared toward accurately separating clumped cells by penalizing foreground voxels when the model confidence is low, which typically occurs near cell membranes.
The three masks generated in the first stage of the pipeline then form the input to the second stage, which performs instance segmentation on the masks. This stage relies on an algorithm called a touching area-based clustering algorithm (Tascan) that separates the 3D cells from the foreground masks.
The second stage does not require any parameters from the first stage, and the Tascan clustering algorithm requires only one hyperparameter (the minimum touching area between two cell foreground super voxels). This makes 3DCellSeg easy to apply to new datasets without the laborious fine-tuning required by previous systems.
Results and future work
The authors compared the segmentation performance of 3DCellSeg against a suite of existing machine-learning systems. The results showed that 3DCellSeg outperformed the baseline models on the ATAS (plant), HMS (animal), and LRP (plant) datasets with an overall accuracy of 95.6%, 76.4%, and 74.7%, respectively. On the Ovules (plant) dataset, 3DCellSeg achieved an overall accuracy of 82.2%, slightly lower than the top-performing U-Net system.
 Comparison of 3DCellSeg against machine-learning systems in ATAS, HMS, and LRP datasets. Image courtesy of Wang et al. Licensed under CC BY 4.0.
Comparison of 3DCellSeg against machine-learning systems in ATAS, HMS, and LRP datasets. Image courtesy of Wang et al. Licensed under CC BY 4.0.Currently, the 3DCellSeg system uses a trained CNN only in the first stage (semantic segmentation). In future work, the researchers hope to harness the power of deep learning alongside the Tascan algorithm in the second stage (clustering) as well. In addition, they hope to incorporate more domain-specific knowledge of the cell characteristics, such as the size and distribution of cells, into the pipeline to improve its accuracy.
The researchers also expect the 3DCellSeg system to be useful as a cell-based disease identification tool, in particular in the area of cancer diagnostics.
"Our novel 3DCellSeg can advance research on 3D instance segmentation; it can serve a powerful cell-based disease identification tool, such as cancer diagnostics, when our cell segmentation model is further trained on labelled human cancer/normal cell images," the authors wrote.